Binding free energy analysis of galectin‐3 natural ligands and synthetic inhibitors
Authors:
Luke Newman, Valerie Vaissier Welborn
Affiliation:
Macromolecules Innovation Institute, Virginia Tech, Blacksburg, Virginia, USA; Department of Chemistry, Virginia Tech, Blacksburg, Virginia, USA
Description:
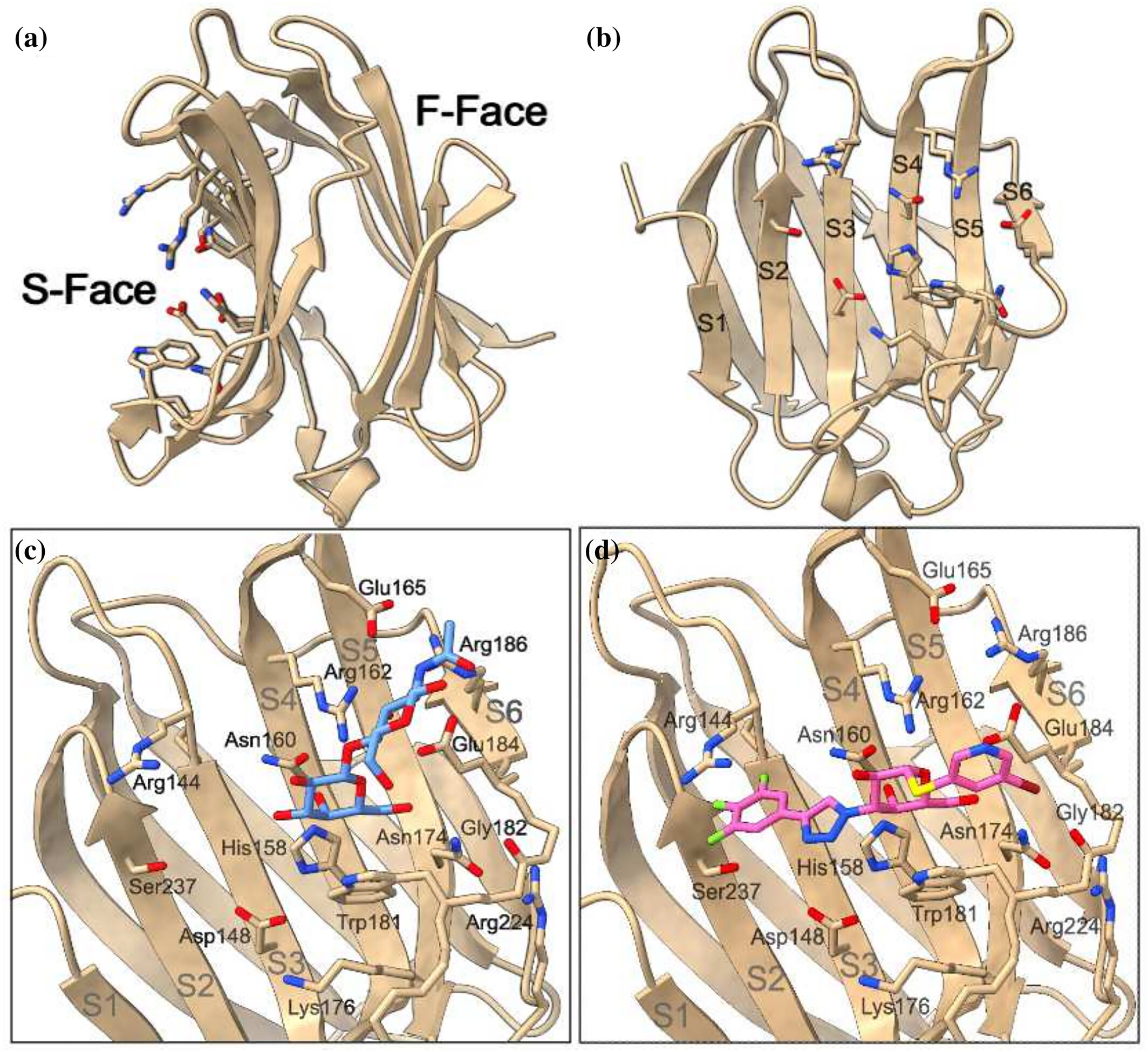
Galectin-3 -ligand complexes are characterized by halogen, σ -hole bonds, hydrogen bonds, cationπ and CHπ interactions. Here, we model these non-covalent interactions with the AMOEBA polarizable force field and conduct an absolute binding free energy analysis on leading galectin-3 inhibitors. Synthetic drug molecules GB0139, GB1107, and GB1211 were estimated to have binding free energies of /C0 4.3, /C0 6.7, and /C0 9.5 kcal/mol respectively. This compares to /C0 0.3 and 1.4 kcal/mol for the natural ligands, N-acetyllactosamine type 1 and type 2, respectively. We calculated the electric fields projected along key bonds in each ligand to further rationalize these results. We find that while the hydroxyl groups of the natural ligands interact reasonably well with residues in galectin-3 ' s binding pocket, structural dynamics weaken the binding pose and favor interactions with water, sometimes yielding to dissociation. In contrast, the more favorable binding energy of GB1211, leading inhibitor in clinical studies, is associated with strong and constant electric fields across the bonds investigated, suggesting a stiffer binding pose with a stabilizing σ -hole interaction.
Publications:
- Luke Newman; Valerie Vaissier Welborn; Binding free energy analysis of galectin‐3 natural ligands and synthetic inhibitors; Protein Science, 2025
Tags:
Atomistic simulation Interaction energies Ligands Modeling and SimulationRelated Chronicles:
Files:
| File Name | File Description | File Type | File Size | File URL |
|---|---|---|---|---|
| Supporting Information | Supporting Information | 3.96 MB | Login to download |