Glycoproteomic characterization of Neuropilin-1 reveals critical glycosylation sites for SARS-CoV-2 entry
Authors:
Tuhin Das, Shuhong Luo, Panning Wang, Jianmin Fang, Asif Shajahan, Lauren Peppi, Sabyasachi Dash, Kino Maravillas, Rochelle N. Wickramasekara, Parastoo Azadi, Ruo-Pan Huang
Affiliation:
RayBiotech Guangzhou Co. Ltd. Guangzhou, Guangzhou 510000, China; RayBiotech Inc, 3607 Parkway Lane, Peachtree Corners, Atlanta, GA 30092, USA; Complex Carbohydrate Research Center, University of Georgia, 315 Riverbend Road, Athens, GA 30602, USA; Center for Vascular Biology, Pathology and Laboratory Medicine, Weill Cornell Medical School. New York, New York 10065, USA
Description:
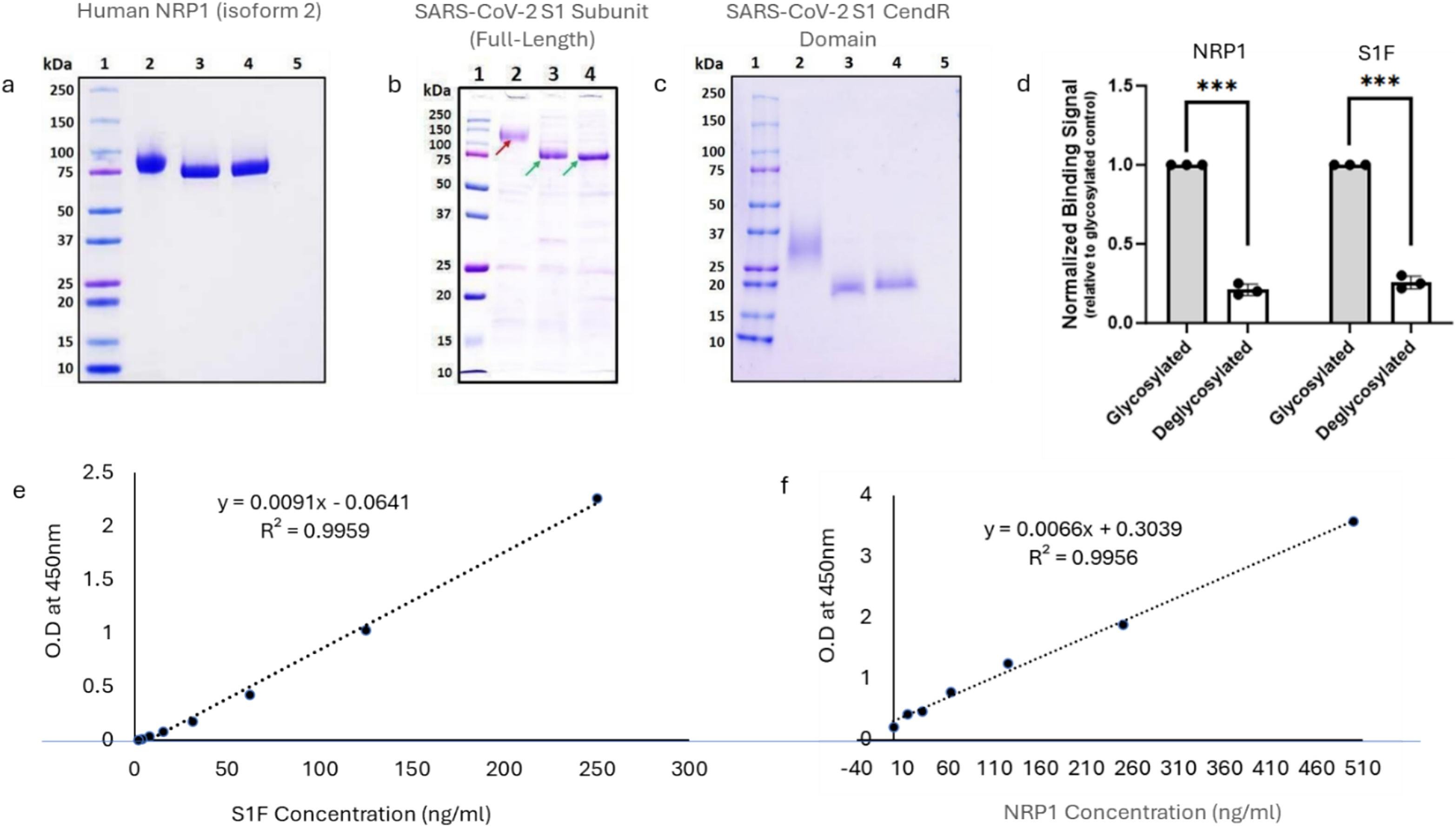
The binding of the SARS‑CoV‑2 spike (S) glycoprotein to human host receptors, including ACE2, NRP1, and AXL, is essential for viral entry. Glycosylation of both the spike protein and its host receptors can significantly influence these interactions. While NRP1 is recognized as a key host receptor, the role of its glycosylation in spike binding and viral infectivity has not been fully elucidated. In this study, we developed a spike‑NRP1 binding assay using recombinant proteins in HEK293 cells and performed LC‑MS‑based glycoproteomic analysis to characterize NRP1 glycosylation. We identified three N‑linked glycosylation sites (N150, N261, N522) and four O‑linked glycosylation sites (S612, S637, T638, S641) on NRP1. Mutational analysis revealed that glycosylation at these specific sites is critical for spike binding, as single‑site mutants showed significantly reduced interaction. Further characterization of terminal sialic acid linkages, bisecting GlcNAc, and fucosylation patterns provided insight into the complexity of NRP1 glycosylation. Functional assays demonstrated that loss of glycosylation impaired SARS‑CoV‑2 pseudovirus entry and altered IL‑6 secretion, indicating a broader role in host immune modulation. These findings demonstrate that site‑specific NRP1 glycosylation modulates spike binding and viral entry in a cell‑based model, providing a foundation for future studies to explore the potential of targeting glycosylation‑dependent mechanisms in SARS‑CoV‑2 infection.
Publications:
- Tuhin Das; Shuhong Luo; Panning Wang; Jianmin Fang; Asif Shajahan; Lauren Peppi; Sabyasachi Dash; Kino Maravillas; Rochelle N. Wickramasekara; Parastoo Azadi; Ruo-Pan Huang; Glycoproteomic characterization of Neuropilin-1 reveals critical glycosylation sites for SARS-CoV-2 entry; Biochimica et Biophysica Acta (BBA) - Proteins and Proteomics, 2025
Tags:
Glycosylation Mass spectrometry Proteomics SARS-CoV-2Files:
| File Name | File Description | File Type | File Size | File URL |
|---|---|---|---|---|
| Supporting Information | Supporting Information | docx | 1.01 MB | Login to download |