Glycosylation of Structured Protein Domains in Cell-Free Reaction Environments
Authors:
Erik J. Bidstrup, Kyle Hill, Chandra K. Bandi, D. Natasha Owitipana, Alina Chisti, Rochelle Aw, Xu Yang, Parastoo Azadi, Michael C. Jewett, Lai-Xi Wang, Weston Kightlinger, Matthew P. DeLisa
Affiliation:
Cell-Free Protein Synthesis and Microbial Process Development, National Resilience Inc., Oakland, California 94606, United States; Department of Chemistry and Biochemistry, University of Maryland, College Park, Maryland 1220742, United States; Complex Carbohydrate Research Center, University of Georgia, Athens, Georgia 30602-4712, United States; Department of Bioengineering, Stanford University, Stanford, California 94305, United States; Robert Frederick Smith School of Chemical and Biomolecular Engineering, Cornell University, Ithaca, New York 14853, United States
Description:
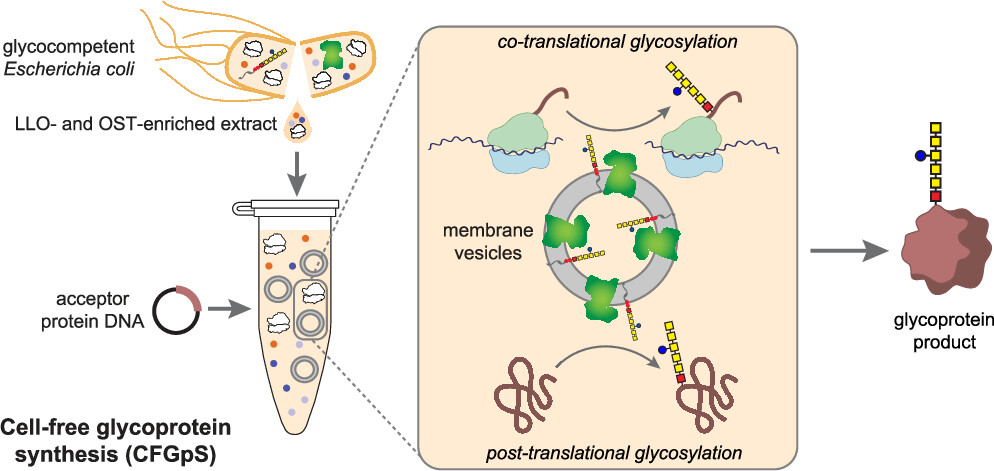
The production of N -linked glycoproteins in genetically tractable bacterial hosts and their cell-free extracts holds great promise for low-cost, customizable, and distributed biomanufacturing of glycoconjugate vaccines and glycoprotein therapeutics. In nearly all bacterial N -linked protein glycosylation systems described so far, a single-subunit, transmembrane oligosaccharyltransferase (OST) is employed which favors acceptor sites in flexible, solvent-exposed motifs of the glycoprotein substrate. Yet despite this preference, acceptor sites in structured domains can also be glycosylated in living bacteria, presumably by a mechanism where the site is presented to the OST in a flexible form during or after the membrane translocation step but prior to
Publications:
- Erik J. Bidstrup; Kyle Hill; Chandra K. Bandi; D. Natasha Owitipana; Alina Chisti; Rochelle Aw; Xu Yang; Parastoo Azadi; Michael C. Jewett; Lai-Xi Wang; Weston Kightlinger; Matthew P. DeLisa; Glycosylation of Structured Protein Domains in Cell-Free Reaction Environments; ACS Synthetic Biology, 2025
Tags:
Carbohydrates Glycosylation Mass spectrometry ProteomicsRelated Chronicles:
Files:
| File Name | File Description | File Type | File Size | File URL |
|---|---|---|---|---|
| Supporting Information | Supporting Information | 9.34 MB | Login to download |